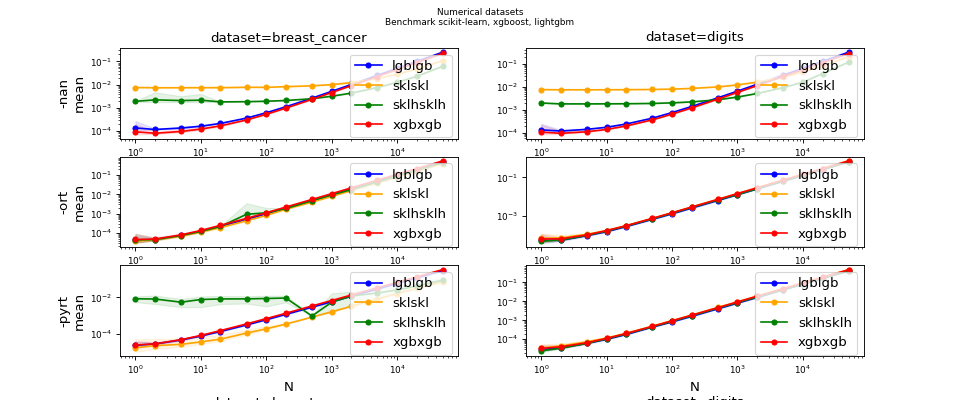
Benchmark (ONNX) for ensemble models#
Overview#
(Source code, png, hires.png, pdf)
Configuration#
<<<
from pyquickhelper.pandashelper import df2rst
import pandas
name = os.path.join(
__WD__, "../../onnx/results/bench_plot_ml_ensemble.time.csv")
df = pandas.read_csv(name)
print(df2rst(df, number_format=4))
>>>
name |
version |
value |
|---|---|---|
date |
2019-12-15 |
|
python |
3.7.2 (default, Mar 1 2019, 18:34:21) [GCC 6.3.0 20170516] |
|
platform |
linux |
|
OS |
Linux-4.9.0-8-amd64-x86_64-with-debian-9.6 |
|
machine |
x86_64 |
|
processor |
||
release |
4.9.0-8-amd64 |
|
architecture |
(‘64bit’, ‘’) |
|
mlprodict |
0.3 |
|
numpy |
1.17.4 |
openblas, language=c |
onnx |
1.6.34 |
opset=12 |
onnxruntime |
1.1.992 |
CPU-DNNL-MKL-ML |
pandas |
0.25.3 |
|
skl2onnx |
1.6.993 |
|
sklearn |
0.22 |
Raw results#
<<<
from pyquickhelper.pandashelper import df2rst
from pymlbenchmark.benchmark.bench_helper import bench_pivot
import pandas
name = os.path.join(
__WD__, "../../onnx/results/bench_plot_ml_ensemble.perf.csv")
df = pandas.read_csv(name)
print(df2rst(df, number_format=4))
lib |
rt |
N |
dataset |
dim |
repeat |
number |
min |
max |
min3 |
max3 |
mean |
lower |
upper |
count |
median |
fit_train_score_.shape |
fit_validation_score_.shape |
fit__predictors.size |
fit__predictors.sum|tree_.leave_count |
fit__predictors.sum|tree_.node_count |
fit__predictors.max|tree_.max_depth |
fit_estimators_.size |
fit_estimators_.sum|tree_.node_count |
fit_estimators_.max|tree_.max_depth |
fit_estimators_.sum|tree_.leave_count |
fit_estimators_.n_features_ |
fit_estimators_.n_classes_ |
fit_objective |
fit_node_count |
fit_ntrees |
fit_leave_count |
fit_mode_count |
fit_n_targets |
|
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
sklh |
1 |
breast_cancer |
-1 |
10 |
1 |
0.001861 |
0.002051 |
0.001889 |
0.001914 |
0.001907 |
0.001861 |
0.002008 |
10 |
0.001892 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
1 |
breast_cancer |
-1 |
10 |
1 |
0.005536 |
0.0113 |
0.008328 |
0.01112 |
0.008437 |
0.005536 |
0.0113 |
10 |
0.008365 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
1 |
breast_cancer |
-1 |
10 |
1 |
3.113e-05 |
0.0001205 |
3.248e-05 |
3.724e-05 |
4.407e-05 |
3.113e-05 |
9.514e-05 |
10 |
3.438e-05 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
2 |
breast_cancer |
-1 |
10 |
1 |
0.001749 |
0.006033 |
0.001759 |
0.001807 |
0.002198 |
0.001749 |
0.004704 |
10 |
0.001764 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
2 |
breast_cancer |
-1 |
10 |
1 |
0.004348 |
0.01282 |
0.008099 |
0.008805 |
0.008202 |
0.004348 |
0.01282 |
10 |
0.00846 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
2 |
breast_cancer |
-1 |
10 |
1 |
4.082e-05 |
5.748e-05 |
4.26e-05 |
4.698e-05 |
4.502e-05 |
4.082e-05 |
5.425e-05 |
10 |
4.281e-05 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
5 |
breast_cancer |
-1 |
10 |
1 |
0.001768 |
0.00323 |
0.001771 |
0.001807 |
0.002061 |
0.001768 |
0.003166 |
10 |
0.001782 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
5 |
breast_cancer |
-1 |
10 |
1 |
0.002943 |
0.008639 |
0.005622 |
0.005894 |
0.005479 |
0.002943 |
0.008236 |
10 |
0.005703 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
5 |
breast_cancer |
-1 |
10 |
1 |
6.981e-05 |
8.635e-05 |
7.163e-05 |
7.236e-05 |
7.323e-05 |
6.981e-05 |
8.21e-05 |
10 |
7.191e-05 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
10 |
breast_cancer |
-1 |
10 |
1 |
0.001766 |
0.00493 |
0.001783 |
0.001937 |
0.002136 |
0.001766 |
0.003966 |
10 |
0.001801 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
10 |
breast_cancer |
-1 |
10 |
1 |
0.002906 |
0.01129 |
0.005827 |
0.01125 |
0.007697 |
0.002906 |
0.01129 |
10 |
0.007192 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
10 |
breast_cancer |
-1 |
10 |
1 |
0.0001151 |
0.0001351 |
0.0001184 |
0.0001218 |
0.0001204 |
0.0001151 |
0.0001309 |
10 |
0.000119 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
20 |
breast_cancer |
-1 |
10 |
1 |
0.001787 |
0.001822 |
0.001793 |
0.001803 |
0.001799 |
0.001787 |
0.00182 |
10 |
0.001798 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
20 |
breast_cancer |
-1 |
10 |
1 |
0.004343 |
0.01208 |
0.008466 |
0.008759 |
0.008256 |
0.004343 |
0.01208 |
10 |
0.008617 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
20 |
breast_cancer |
-1 |
10 |
1 |
0.0002074 |
0.0002321 |
0.0002127 |
0.0002194 |
0.000216 |
0.0002074 |
0.0002295 |
10 |
0.0002136 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
50 |
breast_cancer |
-1 |
10 |
1 |
0.001831 |
0.00187 |
0.001839 |
0.001851 |
0.001845 |
0.001831 |
0.001867 |
10 |
0.001842 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
50 |
breast_cancer |
-1 |
10 |
1 |
0.004623 |
0.01155 |
0.008744 |
0.008912 |
0.008345 |
0.004623 |
0.01155 |
10 |
0.008792 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
50 |
breast_cancer |
-1 |
10 |
1 |
0.0004876 |
0.004791 |
0.0004921 |
0.0004978 |
0.000925 |
0.0004876 |
0.003451 |
10 |
0.0004942 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
100 |
breast_cancer |
-1 |
10 |
1 |
0.001907 |
0.001982 |
0.001913 |
0.001921 |
0.001922 |
0.001907 |
0.001962 |
10 |
0.001915 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
100 |
breast_cancer |
-1 |
10 |
1 |
0.003215 |
0.01246 |
0.007558 |
0.01193 |
0.008725 |
0.003215 |
0.01246 |
10 |
0.009158 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
100 |
breast_cancer |
-1 |
10 |
1 |
0.0009201 |
0.002459 |
0.0009507 |
0.0009707 |
0.001102 |
0.0009201 |
0.001989 |
10 |
0.0009545 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
200 |
breast_cancer |
-1 |
10 |
1 |
0.002051 |
0.00212 |
0.002061 |
0.002084 |
0.002074 |
0.002051 |
0.00212 |
10 |
0.002066 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
200 |
breast_cancer |
-1 |
10 |
1 |
0.005421 |
0.01272 |
0.009619 |
0.009825 |
0.009262 |
0.005421 |
0.01272 |
10 |
0.009682 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
200 |
breast_cancer |
-1 |
10 |
1 |
0.001817 |
0.001918 |
0.001828 |
0.001853 |
0.001846 |
0.001817 |
0.001907 |
10 |
0.001833 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
500 |
breast_cancer |
-1 |
10 |
1 |
0.002442 |
0.002515 |
0.002456 |
0.002473 |
0.002465 |
0.002442 |
0.002504 |
10 |
0.002464 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
500 |
breast_cancer |
-1 |
10 |
1 |
0.0009548 |
0.0009952 |
0.0009567 |
0.0009636 |
0.0009623 |
0.0009548 |
0.0009848 |
10 |
0.0009582 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
500 |
breast_cancer |
-1 |
10 |
1 |
0.004649 |
0.004712 |
0.004661 |
0.004674 |
0.00467 |
0.004649 |
0.004702 |
10 |
0.004665 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
1000 |
breast_cancer |
-1 |
10 |
1 |
0.003094 |
0.003536 |
0.003116 |
0.003136 |
0.00316 |
0.003094 |
0.003408 |
10 |
0.00312 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
1000 |
breast_cancer |
-1 |
10 |
1 |
0.001481 |
0.01687 |
0.001893 |
0.008945 |
0.005436 |
0.001481 |
0.01687 |
10 |
0.0019 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
1000 |
breast_cancer |
-1 |
10 |
1 |
0.008786 |
0.01228 |
0.009021 |
0.009297 |
0.009428 |
0.008786 |
0.01133 |
10 |
0.009222 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
2000 |
breast_cancer |
-1 |
10 |
1 |
0.004283 |
0.004444 |
0.004286 |
0.0043 |
0.004307 |
0.004283 |
0.004398 |
10 |
0.004293 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
2000 |
breast_cancer |
-1 |
10 |
1 |
0.003786 |
0.02055 |
0.003836 |
0.02024 |
0.0119 |
0.003786 |
0.02055 |
10 |
0.0119 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
2000 |
breast_cancer |
-1 |
10 |
1 |
0.01744 |
0.01796 |
0.01749 |
0.01762 |
0.01758 |
0.01744 |
0.01786 |
10 |
0.01756 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
5000 |
breast_cancer |
-1 |
10 |
1 |
0.007426 |
0.007676 |
0.007448 |
0.007471 |
0.007487 |
0.007426 |
0.00764 |
10 |
0.007453 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
5000 |
breast_cancer |
-1 |
10 |
1 |
0.009353 |
0.03219 |
0.009511 |
0.0258 |
0.0175 |
0.009353 |
0.03219 |
10 |
0.01598 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
5000 |
breast_cancer |
-1 |
10 |
1 |
0.04369 |
0.04592 |
0.04385 |
0.04456 |
0.04443 |
0.04369 |
0.04592 |
10 |
0.04422 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
10000 |
breast_cancer |
-1 |
10 |
1 |
0.01252 |
0.01268 |
0.01253 |
0.01257 |
0.01256 |
0.01252 |
0.01266 |
10 |
0.01255 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
10000 |
breast_cancer |
-1 |
10 |
1 |
0.01888 |
0.03861 |
0.01893 |
0.03227 |
0.02562 |
0.01888 |
0.03861 |
10 |
0.02264 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
10000 |
breast_cancer |
-1 |
10 |
1 |
0.08726 |
0.08811 |
0.08739 |
0.08784 |
0.08755 |
0.08726 |
0.08808 |
10 |
0.08741 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
20000 |
breast_cancer |
-1 |
10 |
1 |
0.02268 |
0.02301 |
0.02276 |
0.02285 |
0.02281 |
0.02268 |
0.02301 |
10 |
0.02279 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
20000 |
breast_cancer |
-1 |
10 |
1 |
0.03303 |
0.07256 |
0.03646 |
0.04025 |
0.04211 |
0.03303 |
0.06614 |
10 |
0.0368 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
20000 |
breast_cancer |
-1 |
10 |
1 |
0.1749 |
0.1762 |
0.1751 |
0.1755 |
0.1753 |
0.1749 |
0.1761 |
10 |
0.1752 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
50000 |
breast_cancer |
-1 |
10 |
1 |
0.06184 |
0.06285 |
0.06221 |
0.06233 |
0.06227 |
0.06184 |
0.06277 |
10 |
0.06227 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
|||||||||||||
sklh |
pyrt |
50000 |
breast_cancer |
-1 |
10 |
1 |
0.07937 |
0.1379 |
0.08035 |
0.08856 |
0.08917 |
0.07937 |
0.1227 |
10 |
0.08115 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
sklh |
ort |
50000 |
breast_cancer |
-1 |
10 |
1 |
0.4375 |
0.4441 |
0.4385 |
0.4418 |
0.44 |
0.4375 |
0.4441 |
10 |
0.4388 |
0 |
0 |
30 |
100 |
1160 |
2220 |
6 |
||||||||||||
skl |
1 |
breast_cancer |
-1 |
10 |
1 |
0.007443 |
0.007781 |
0.007482 |
0.007509 |
0.007516 |
0.007443 |
0.007696 |
10 |
0.007496 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
1 |
breast_cancer |
-1 |
10 |
1 |
1.039e-05 |
6.953e-05 |
1.097e-05 |
1.271e-05 |
1.732e-05 |
1.039e-05 |
5.152e-05 |
10 |
1.109e-05 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
1 |
breast_cancer |
-1 |
10 |
1 |
2.948e-05 |
8.007e-05 |
3.009e-05 |
3.455e-05 |
3.689e-05 |
2.948e-05 |
6.59e-05 |
10 |
3.108e-05 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
2 |
breast_cancer |
-1 |
10 |
1 |
0.007295 |
0.007445 |
0.007314 |
0.007361 |
0.007346 |
0.007295 |
0.007445 |
10 |
0.007321 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
2 |
breast_cancer |
-1 |
10 |
1 |
1.501e-05 |
6.764e-05 |
1.721e-05 |
1.957e-05 |
2.28e-05 |
1.501e-05 |
5.239e-05 |
10 |
1.782e-05 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
2 |
breast_cancer |
-1 |
10 |
1 |
3.935e-05 |
5.732e-05 |
4.059e-05 |
4.302e-05 |
4.333e-05 |
3.935e-05 |
5.341e-05 |
10 |
4.128e-05 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
5 |
breast_cancer |
-1 |
10 |
1 |
0.00733 |
0.007482 |
0.007338 |
0.00736 |
0.007362 |
0.00733 |
0.007447 |
10 |
0.007348 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
5 |
breast_cancer |
-1 |
10 |
1 |
1.934e-05 |
6.705e-05 |
2.225e-05 |
2.428e-05 |
2.694e-05 |
1.934e-05 |
5.336e-05 |
10 |
2.301e-05 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
5 |
breast_cancer |
-1 |
10 |
1 |
6.115e-05 |
7.96e-05 |
6.486e-05 |
7.1e-05 |
6.751e-05 |
6.115e-05 |
7.742e-05 |
10 |
6.586e-05 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
10 |
breast_cancer |
-1 |
10 |
1 |
0.007351 |
0.007596 |
0.007364 |
0.007379 |
0.007396 |
0.007351 |
0.007533 |
10 |
0.00737 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
10 |
breast_cancer |
-1 |
10 |
1 |
2.763e-05 |
8.745e-05 |
2.902e-05 |
3.165e-05 |
3.641e-05 |
2.763e-05 |
7.036e-05 |
10 |
3.053e-05 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
10 |
breast_cancer |
-1 |
10 |
1 |
0.0001022 |
0.0001235 |
0.0001064 |
0.0001132 |
0.0001101 |
0.0001022 |
0.0001216 |
10 |
0.0001081 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
20 |
breast_cancer |
-1 |
10 |
1 |
0.007356 |
0.007514 |
0.007373 |
0.007418 |
0.007404 |
0.007356 |
0.007511 |
10 |
0.007378 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
20 |
breast_cancer |
-1 |
10 |
1 |
4.13e-05 |
0.0001102 |
4.37e-05 |
4.713e-05 |
5.168e-05 |
4.13e-05 |
9.036e-05 |
10 |
4.492e-05 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
20 |
breast_cancer |
-1 |
10 |
1 |
0.0001745 |
0.0002095 |
0.0001825 |
0.0001984 |
0.0001895 |
0.0001745 |
0.0002095 |
10 |
0.0001887 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
50 |
breast_cancer |
-1 |
10 |
1 |
0.007515 |
0.008537 |
0.007526 |
0.007593 |
0.007689 |
0.007515 |
0.008307 |
10 |
0.00755 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
50 |
breast_cancer |
-1 |
10 |
1 |
9.159e-05 |
0.0001965 |
9.966e-05 |
0.0001087 |
0.0001113 |
9.159e-05 |
0.0001678 |
10 |
0.0001027 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
50 |
breast_cancer |
-1 |
10 |
1 |
0.0003977 |
0.0005152 |
0.0004104 |
0.0004402 |
0.0004278 |
0.0003977 |
0.0004903 |
10 |
0.0004149 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
100 |
breast_cancer |
-1 |
10 |
1 |
0.00758 |
0.007754 |
0.007613 |
0.007624 |
0.007636 |
0.00758 |
0.007734 |
10 |
0.00762 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
100 |
breast_cancer |
-1 |
10 |
1 |
0.0001721 |
0.0003097 |
0.000174 |
0.0001804 |
0.000191 |
0.0001721 |
0.0002695 |
10 |
0.0001771 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
100 |
breast_cancer |
-1 |
10 |
1 |
0.0007968 |
0.0008729 |
0.0008006 |
0.000808 |
0.0008117 |
0.0007968 |
0.0008562 |
10 |
0.0008024 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
200 |
breast_cancer |
-1 |
10 |
1 |
0.00802 |
0.008874 |
0.008045 |
0.0082 |
0.008187 |
0.00802 |
0.008683 |
10 |
0.008062 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
200 |
breast_cancer |
-1 |
10 |
1 |
0.0003351 |
0.0004204 |
0.0003361 |
0.0003432 |
0.0003463 |
0.0003351 |
0.0003951 |
10 |
0.0003373 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
200 |
breast_cancer |
-1 |
10 |
1 |
0.001597 |
0.001733 |
0.001697 |
0.001728 |
0.001696 |
0.00162 |
0.001733 |
10 |
0.001701 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
500 |
breast_cancer |
-1 |
10 |
1 |
0.00874 |
0.009575 |
0.008782 |
0.00888 |
0.008896 |
0.00874 |
0.009355 |
10 |
0.008816 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
500 |
breast_cancer |
-1 |
10 |
1 |
0.0008108 |
0.0009033 |
0.0008222 |
0.0008263 |
0.0008301 |
0.0008108 |
0.0008791 |
10 |
0.0008226 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
500 |
breast_cancer |
-1 |
10 |
1 |
0.003879 |
0.00404 |
0.003903 |
0.003941 |
0.003926 |
0.003879 |
0.004013 |
10 |
0.003914 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
1000 |
breast_cancer |
-1 |
10 |
1 |
0.009885 |
0.01017 |
0.009962 |
0.009998 |
0.009982 |
0.009885 |
0.01013 |
10 |
0.009974 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
1000 |
breast_cancer |
-1 |
10 |
1 |
0.001614 |
0.001772 |
0.001624 |
0.001636 |
0.001641 |
0.001614 |
0.001728 |
10 |
0.001627 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
1000 |
breast_cancer |
-1 |
10 |
1 |
0.007717 |
0.00793 |
0.007771 |
0.007801 |
0.007796 |
0.007717 |
0.007899 |
10 |
0.007797 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
2000 |
breast_cancer |
-1 |
10 |
1 |
0.01204 |
0.01237 |
0.01214 |
0.01225 |
0.01218 |
0.01204 |
0.01236 |
10 |
0.01216 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
2000 |
breast_cancer |
-1 |
10 |
1 |
0.003202 |
0.003326 |
0.003229 |
0.003246 |
0.003246 |
0.003202 |
0.003322 |
10 |
0.003233 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
2000 |
breast_cancer |
-1 |
10 |
1 |
0.01545 |
0.01614 |
0.01554 |
0.01561 |
0.01562 |
0.01545 |
0.01598 |
10 |
0.01555 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
5000 |
breast_cancer |
-1 |
10 |
1 |
0.01814 |
0.01908 |
0.01827 |
0.01836 |
0.01837 |
0.01814 |
0.01887 |
10 |
0.01828 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
5000 |
breast_cancer |
-1 |
10 |
1 |
0.007987 |
0.009007 |
0.008083 |
0.008129 |
0.008172 |
0.007987 |
0.008725 |
10 |
0.008095 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
5000 |
breast_cancer |
-1 |
10 |
1 |
0.0387 |
0.03954 |
0.03885 |
0.03912 |
0.03896 |
0.0387 |
0.03944 |
10 |
0.0389 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
10000 |
breast_cancer |
-1 |
10 |
1 |
0.02782 |
0.02829 |
0.02792 |
0.02806 |
0.02799 |
0.02782 |
0.02829 |
10 |
0.02798 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
10000 |
breast_cancer |
-1 |
10 |
1 |
0.01604 |
0.01618 |
0.01609 |
0.01614 |
0.0161 |
0.01604 |
0.01618 |
10 |
0.0161 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
10000 |
breast_cancer |
-1 |
10 |
1 |
0.0774 |
0.07822 |
0.07752 |
0.07772 |
0.07763 |
0.0774 |
0.07808 |
10 |
0.07756 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
20000 |
breast_cancer |
-1 |
10 |
1 |
0.04701 |
0.0475 |
0.04724 |
0.0473 |
0.04726 |
0.04704 |
0.04749 |
10 |
0.04728 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
20000 |
breast_cancer |
-1 |
10 |
1 |
0.02691 |
0.03227 |
0.03194 |
0.03211 |
0.03147 |
0.02845 |
0.03227 |
10 |
0.03199 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
20000 |
breast_cancer |
-1 |
10 |
1 |
0.1549 |
0.1557 |
0.1551 |
0.1555 |
0.1553 |
0.1549 |
0.1557 |
10 |
0.1552 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
50000 |
breast_cancer |
-1 |
10 |
1 |
0.1069 |
0.1112 |
0.107 |
0.1083 |
0.1079 |
0.1069 |
0.1105 |
10 |
0.1071 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
|||||||||||||
skl |
pyrt |
50000 |
breast_cancer |
-1 |
10 |
1 |
0.06958 |
0.07694 |
0.07058 |
0.07345 |
0.07218 |
0.06958 |
0.07694 |
10 |
0.07143 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
skl |
ort |
50000 |
breast_cancer |
-1 |
10 |
1 |
0.3877 |
0.3912 |
0.3882 |
0.3889 |
0.3888 |
0.3877 |
0.3906 |
10 |
0.3885 |
30 |
100 |
2466 |
6 |
1283 |
30 |
1 |
||||||||||||
xgb |
1 |
breast_cancer |
-1 |
10 |
1 |
7.096e-05 |
0.0002602 |
7.199e-05 |
7.601e-05 |
9.248e-05 |
7.096e-05 |
0.0002023 |
10 |
7.236e-05 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
1 |
breast_cancer |
-1 |
10 |
1 |
1.768e-05 |
4.082e-05 |
2.03e-05 |
2.586e-05 |
2.486e-05 |
1.768e-05 |
3.93e-05 |
10 |
2.195e-05 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
1 |
breast_cancer |
-1 |
10 |
1 |
3.143e-05 |
0.0001099 |
3.632e-05 |
4.301e-05 |
4.573e-05 |
3.143e-05 |
8.923e-05 |
10 |
3.741e-05 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
2 |
breast_cancer |
-1 |
10 |
1 |
7.503e-05 |
9.641e-05 |
7.667e-05 |
8.14e-05 |
8.002e-05 |
7.503e-05 |
9.233e-05 |
10 |
7.807e-05 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
2 |
breast_cancer |
-1 |
10 |
1 |
2.528e-05 |
3.82e-05 |
2.65e-05 |
2.833e-05 |
2.853e-05 |
2.528e-05 |
3.573e-05 |
10 |
2.734e-05 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
2 |
breast_cancer |
-1 |
10 |
1 |
4.194e-05 |
6.383e-05 |
4.534e-05 |
4.912e-05 |
4.83e-05 |
4.194e-05 |
6.002e-05 |
10 |
4.647e-05 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
5 |
breast_cancer |
-1 |
10 |
1 |
8.759e-05 |
0.0001238 |
8.904e-05 |
9.36e-05 |
9.546e-05 |
8.759e-05 |
0.0001173 |
10 |
9.076e-05 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
5 |
breast_cancer |
-1 |
10 |
1 |
4.263e-05 |
5.848e-05 |
4.762e-05 |
4.857e-05 |
4.86e-05 |
4.263e-05 |
5.634e-05 |
10 |
4.765e-05 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
5 |
breast_cancer |
-1 |
10 |
1 |
7.177e-05 |
9.445e-05 |
7.758e-05 |
7.889e-05 |
7.964e-05 |
7.177e-05 |
9.136e-05 |
10 |
7.837e-05 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
10 |
breast_cancer |
-1 |
10 |
1 |
0.0001103 |
0.0001401 |
0.0001151 |
0.0001276 |
0.0001212 |
0.0001103 |
0.0001401 |
10 |
0.0001166 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
10 |
breast_cancer |
-1 |
10 |
1 |
7.671e-05 |
9.357e-05 |
8.1e-05 |
8.45e-05 |
8.283e-05 |
7.671e-05 |
9.207e-05 |
10 |
8.13e-05 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
10 |
breast_cancer |
-1 |
10 |
1 |
0.0001235 |
0.0001513 |
0.0001318 |
0.0001345 |
0.0001341 |
0.0001235 |
0.0001485 |
10 |
0.0001322 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
20 |
breast_cancer |
-1 |
10 |
1 |
0.000156 |
0.0001897 |
0.0001616 |
0.0001693 |
0.0001669 |
0.000156 |
0.0001883 |
10 |
0.0001624 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
20 |
breast_cancer |
-1 |
10 |
1 |
0.0001503 |
0.0001628 |
0.0001514 |
0.0001536 |
0.0001532 |
0.0001503 |
0.0001605 |
10 |
0.0001517 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
20 |
breast_cancer |
-1 |
10 |
1 |
0.00024 |
0.0002726 |
0.0002439 |
0.0002503 |
0.000249 |
0.00024 |
0.0002688 |
10 |
0.0002445 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
50 |
breast_cancer |
-1 |
10 |
1 |
0.0002905 |
0.0003191 |
0.0002951 |
0.0003067 |
0.0003001 |
0.0002905 |
0.0003171 |
10 |
0.0002988 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
50 |
breast_cancer |
-1 |
10 |
1 |
0.0003462 |
0.0003691 |
0.0003512 |
0.0003583 |
0.0003549 |
0.0003462 |
0.000367 |
10 |
0.0003528 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
50 |
breast_cancer |
-1 |
10 |
1 |
0.000552 |
0.0005867 |
0.0005652 |
0.0005708 |
0.0005677 |
0.000552 |
0.0005844 |
10 |
0.0005668 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
100 |
breast_cancer |
-1 |
10 |
1 |
0.0005115 |
0.0005505 |
0.0005152 |
0.0005376 |
0.0005252 |
0.0005115 |
0.0005505 |
10 |
0.0005202 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
100 |
breast_cancer |
-1 |
10 |
1 |
0.0006773 |
0.0007076 |
0.000683 |
0.0006954 |
0.0006899 |
0.0006773 |
0.0007076 |
10 |
0.0006907 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
100 |
breast_cancer |
-1 |
10 |
1 |
0.001091 |
0.001134 |
0.001108 |
0.00112 |
0.00111 |
0.001091 |
0.001134 |
10 |
0.00111 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
200 |
breast_cancer |
-1 |
10 |
1 |
0.0009517 |
0.001017 |
0.0009641 |
0.0009738 |
0.0009716 |
0.0009517 |
0.001004 |
10 |
0.0009667 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
200 |
breast_cancer |
-1 |
10 |
1 |
0.001353 |
0.001385 |
0.001361 |
0.001367 |
0.001364 |
0.001353 |
0.001381 |
10 |
0.001363 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
200 |
breast_cancer |
-1 |
10 |
1 |
0.002171 |
0.002219 |
0.002184 |
0.002213 |
0.002195 |
0.002171 |
0.002219 |
10 |
0.002192 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
500 |
breast_cancer |
-1 |
10 |
1 |
0.002297 |
0.002336 |
0.002306 |
0.00232 |
0.002311 |
0.002297 |
0.002333 |
10 |
0.00231 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
500 |
breast_cancer |
-1 |
10 |
1 |
0.003377 |
0.003527 |
0.003397 |
0.003417 |
0.003419 |
0.003377 |
0.003502 |
10 |
0.003403 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
500 |
breast_cancer |
-1 |
10 |
1 |
0.005406 |
0.005511 |
0.00543 |
0.005447 |
0.005444 |
0.005406 |
0.005504 |
10 |
0.005441 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
1000 |
breast_cancer |
-1 |
10 |
1 |
0.004525 |
0.00461 |
0.004539 |
0.004565 |
0.004551 |
0.004525 |
0.004598 |
10 |
0.004547 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
1000 |
breast_cancer |
-1 |
10 |
1 |
0.006734 |
0.006817 |
0.00676 |
0.006797 |
0.006776 |
0.006734 |
0.006817 |
10 |
0.006773 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
1000 |
breast_cancer |
-1 |
10 |
1 |
0.01078 |
0.0109 |
0.01084 |
0.01089 |
0.01086 |
0.01078 |
0.0109 |
10 |
0.01087 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
2000 |
breast_cancer |
-1 |
10 |
1 |
0.008969 |
0.009069 |
0.009012 |
0.009054 |
0.009026 |
0.008969 |
0.009069 |
10 |
0.009027 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
2000 |
breast_cancer |
-1 |
10 |
1 |
0.01342 |
0.01375 |
0.0135 |
0.01361 |
0.01354 |
0.01342 |
0.01372 |
10 |
0.01351 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
2000 |
breast_cancer |
-1 |
10 |
1 |
0.02157 |
0.02183 |
0.02164 |
0.02171 |
0.02167 |
0.02157 |
0.02183 |
10 |
0.02166 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
5000 |
breast_cancer |
-1 |
10 |
1 |
0.02237 |
0.02245 |
0.02241 |
0.02244 |
0.02242 |
0.02237 |
0.02245 |
10 |
0.02241 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
5000 |
breast_cancer |
-1 |
10 |
1 |
0.03362 |
0.03439 |
0.03373 |
0.03379 |
0.03382 |
0.03362 |
0.03421 |
10 |
0.03377 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
5000 |
breast_cancer |
-1 |
10 |
1 |
0.05402 |
0.05437 |
0.05412 |
0.05417 |
0.05416 |
0.05402 |
0.05435 |
10 |
0.05415 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
10000 |
breast_cancer |
-1 |
10 |
1 |
0.04462 |
0.04505 |
0.04471 |
0.04478 |
0.04477 |
0.04462 |
0.04498 |
10 |
0.04474 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
10000 |
breast_cancer |
-1 |
10 |
1 |
0.06737 |
0.06779 |
0.06746 |
0.06754 |
0.06751 |
0.06737 |
0.06773 |
10 |
0.0675 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
10000 |
breast_cancer |
-1 |
10 |
1 |
0.1082 |
0.1095 |
0.1084 |
0.1089 |
0.1086 |
0.1082 |
0.1094 |
10 |
0.1086 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
20000 |
breast_cancer |
-1 |
10 |
1 |
0.08925 |
0.0896 |
0.08937 |
0.08944 |
0.0894 |
0.08925 |
0.08958 |
10 |
0.08941 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
20000 |
breast_cancer |
-1 |
10 |
1 |
0.1348 |
0.1361 |
0.135 |
0.1354 |
0.1352 |
0.1348 |
0.1359 |
10 |
0.1351 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
20000 |
breast_cancer |
-1 |
10 |
1 |
0.2161 |
0.2173 |
0.2168 |
0.2169 |
0.2168 |
0.2163 |
0.2173 |
10 |
0.2168 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
50000 |
breast_cancer |
-1 |
10 |
1 |
0.2232 |
0.2237 |
0.2233 |
0.2236 |
0.2234 |
0.2232 |
0.2237 |
10 |
0.2235 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
||||||||||||||
xgb |
pyrt |
50000 |
breast_cancer |
-1 |
10 |
1 |
0.337 |
0.3377 |
0.3374 |
0.3375 |
0.3374 |
0.337 |
0.3377 |
10 |
0.3374 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
xgb |
ort |
50000 |
breast_cancer |
-1 |
10 |
1 |
0.5404 |
0.5452 |
0.5411 |
0.5416 |
0.5417 |
0.5404 |
0.5442 |
10 |
0.5413 |
100 |
reg:linear |
4582 |
100 |
2341 |
2 |
|||||||||||||
lgb |
1 |
breast_cancer |
-1 |
10 |
1 |
0.0001066 |
0.0003401 |
0.0001086 |
0.0001182 |
0.0001359 |
0.0001066 |
0.0002701 |
10 |
0.0001118 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
1 |
breast_cancer |
-1 |
10 |
1 |
1.91e-05 |
4.617e-05 |
2.045e-05 |
2.387e-05 |
2.399e-05 |
1.91e-05 |
3.91e-05 |
10 |
2.058e-05 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
1 |
breast_cancer |
-1 |
10 |
1 |
3.128e-05 |
9.415e-05 |
3.315e-05 |
3.807e-05 |
4.074e-05 |
3.128e-05 |
7.653e-05 |
10 |
3.359e-05 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
2 |
breast_cancer |
-1 |
10 |
1 |
0.000111 |
0.0001312 |
0.0001125 |
0.0001189 |
0.0001164 |
0.000111 |
0.000128 |
10 |
0.0001143 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
2 |
breast_cancer |
-1 |
10 |
1 |
2.612e-05 |
3.542e-05 |
2.671e-05 |
2.826e-05 |
2.808e-05 |
2.612e-05 |
3.329e-05 |
10 |
2.722e-05 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
2 |
breast_cancer |
-1 |
10 |
1 |
4.161e-05 |
5.86e-05 |
4.217e-05 |
4.543e-05 |
4.501e-05 |
4.161e-05 |
5.466e-05 |
10 |
4.309e-05 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
5 |
breast_cancer |
-1 |
10 |
1 |
0.0001264 |
0.0001571 |
0.0001292 |
0.0001378 |
0.0001346 |
0.0001264 |
0.0001528 |
10 |
0.0001313 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
5 |
breast_cancer |
-1 |
10 |
1 |
4.436e-05 |
5.503e-05 |
4.546e-05 |
4.72e-05 |
4.676e-05 |
4.436e-05 |
5.254e-05 |
10 |
4.555e-05 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
5 |
breast_cancer |
-1 |
10 |
1 |
7.075e-05 |
8.758e-05 |
7.194e-05 |
7.332e-05 |
7.399e-05 |
7.075e-05 |
8.322e-05 |
10 |
7.23e-05 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
10 |
breast_cancer |
-1 |
10 |
1 |
0.0001533 |
0.0001785 |
0.0001579 |
0.0001639 |
0.000161 |
0.0001533 |
0.0001755 |
10 |
0.0001584 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
10 |
breast_cancer |
-1 |
10 |
1 |
7.399e-05 |
8.467e-05 |
7.458e-05 |
7.69e-05 |
7.656e-05 |
7.399e-05 |
8.278e-05 |
10 |
7.573e-05 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
10 |
breast_cancer |
-1 |
10 |
1 |
0.0001171 |
0.0001355 |
0.0001185 |
0.0001215 |
0.0001212 |
0.0001171 |
0.0001314 |
10 |
0.0001191 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
20 |
breast_cancer |
-1 |
10 |
1 |
0.0002049 |
0.0002352 |
0.0002078 |
0.0002138 |
0.0002128 |
0.0002049 |
0.0002307 |
10 |
0.0002089 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
20 |
breast_cancer |
-1 |
10 |
1 |
0.000132 |
0.0001446 |
0.0001346 |
0.0001356 |
0.0001357 |
0.000132 |
0.0001424 |
10 |
0.0001351 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
20 |
breast_cancer |
-1 |
10 |
1 |
0.0002094 |
0.000229 |
0.0002131 |
0.0002174 |
0.0002156 |
0.0002094 |
0.0002263 |
10 |
0.0002141 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
50 |
breast_cancer |
-1 |
10 |
1 |
0.0003542 |
0.0003777 |
0.0003552 |
0.0003615 |
0.0003602 |
0.0003542 |
0.0003732 |
10 |
0.0003595 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
50 |
breast_cancer |
-1 |
10 |
1 |
0.0003037 |
0.0003231 |
0.0003072 |
0.000309 |
0.0003089 |
0.0003037 |
0.0003191 |
10 |
0.0003078 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
50 |
breast_cancer |
-1 |
10 |
1 |
0.0004844 |
0.0005103 |
0.000489 |
0.0004921 |
0.0004919 |
0.0004844 |
0.000506 |
10 |
0.0004899 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
100 |
breast_cancer |
-1 |
10 |
1 |
0.0006049 |
0.0006304 |
0.0006098 |
0.0006171 |
0.0006132 |
0.0006049 |
0.0006269 |
10 |
0.0006117 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
100 |
breast_cancer |
-1 |
10 |
1 |
0.0005927 |
0.0006165 |
0.0005964 |
0.0005992 |
0.000599 |
0.0005927 |
0.0006116 |
10 |
0.000597 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
100 |
breast_cancer |
-1 |
10 |
1 |
0.0009486 |
0.001009 |
0.0009493 |
0.0009601 |
0.0009605 |
0.0009486 |
0.0009956 |
10 |
0.0009538 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
200 |
breast_cancer |
-1 |
10 |
1 |
0.001101 |
0.001129 |
0.001108 |
0.001124 |
0.001115 |
0.001101 |
0.001129 |
10 |
0.001113 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
200 |
breast_cancer |
-1 |
10 |
1 |
0.001159 |
0.00121 |
0.001171 |
0.001186 |
0.00118 |
0.001159 |
0.00121 |
10 |
0.001174 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
200 |
breast_cancer |
-1 |
10 |
1 |
0.001857 |
0.001993 |
0.001882 |
0.00192 |
0.001903 |
0.001857 |
0.001986 |
10 |
0.00189 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
500 |
breast_cancer |
-1 |
10 |
1 |
0.002588 |
0.002625 |
0.002615 |
0.002619 |
0.002613 |
0.002592 |
0.002625 |
10 |
0.002616 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
500 |
breast_cancer |
-1 |
10 |
1 |
0.002886 |
0.002936 |
0.002907 |
0.002916 |
0.002909 |
0.002886 |
0.002934 |
10 |
0.002907 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
500 |
breast_cancer |
-1 |
10 |
1 |
0.004617 |
0.004685 |
0.004631 |
0.004657 |
0.004644 |
0.004617 |
0.004685 |
10 |
0.004637 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
1000 |
breast_cancer |
-1 |
10 |
1 |
0.005113 |
0.005194 |
0.005123 |
0.005161 |
0.005142 |
0.005113 |
0.005189 |
10 |
0.005141 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
1000 |
breast_cancer |
-1 |
10 |
1 |
0.00578 |
0.005876 |
0.005789 |
0.005834 |
0.005811 |
0.00578 |
0.005869 |
10 |
0.005806 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
1000 |
breast_cancer |
-1 |
10 |
1 |
0.00922 |
0.009354 |
0.009235 |
0.009258 |
0.009259 |
0.00922 |
0.009334 |
10 |
0.009247 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
2000 |
breast_cancer |
-1 |
10 |
1 |
0.01012 |
0.01019 |
0.01013 |
0.01017 |
0.01015 |
0.01012 |
0.01019 |
10 |
0.01015 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
2000 |
breast_cancer |
-1 |
10 |
1 |
0.01155 |
0.01164 |
0.01156 |
0.01157 |
0.01157 |
0.01155 |
0.01162 |
10 |
0.01156 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
2000 |
breast_cancer |
-1 |
10 |
1 |
0.01841 |
0.0185 |
0.01843 |
0.01849 |
0.01845 |
0.01841 |
0.0185 |
10 |
0.01844 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
5000 |
breast_cancer |
-1 |
10 |
1 |
0.02509 |
0.02528 |
0.0252 |
0.02525 |
0.02521 |
0.0251 |
0.02528 |
10 |
0.02522 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
5000 |
breast_cancer |
-1 |
10 |
1 |
0.02884 |
0.02907 |
0.02888 |
0.02898 |
0.02893 |
0.02884 |
0.02906 |
10 |
0.02893 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
5000 |
breast_cancer |
-1 |
10 |
1 |
0.04596 |
0.04627 |
0.046 |
0.0462 |
0.04608 |
0.04596 |
0.04627 |
10 |
0.04603 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
10000 |
breast_cancer |
-1 |
10 |
1 |
0.05028 |
0.05055 |
0.05037 |
0.05045 |
0.05042 |
0.05028 |
0.05055 |
10 |
0.05042 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
10000 |
breast_cancer |
-1 |
10 |
1 |
0.05772 |
0.05797 |
0.05776 |
0.05787 |
0.0578 |
0.05772 |
0.05796 |
10 |
0.05777 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
10000 |
breast_cancer |
-1 |
10 |
1 |
0.09193 |
0.09267 |
0.09202 |
0.09212 |
0.09214 |
0.09193 |
0.09259 |
10 |
0.09208 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
20000 |
breast_cancer |
-1 |
10 |
1 |
0.1005 |
0.1014 |
0.1006 |
0.1009 |
0.1008 |
0.1005 |
0.1013 |
10 |
0.1007 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
20000 |
breast_cancer |
-1 |
10 |
1 |
0.1155 |
0.1158 |
0.1155 |
0.1156 |
0.1156 |
0.1155 |
0.1158 |
10 |
0.1156 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
20000 |
breast_cancer |
-1 |
10 |
1 |
0.1839 |
0.1846 |
0.1841 |
0.1842 |
0.1841 |
0.1839 |
0.1845 |
10 |
0.1841 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
50000 |
breast_cancer |
-1 |
10 |
1 |
0.251 |
0.2526 |
0.2515 |
0.2523 |
0.2518 |
0.251 |
0.2526 |
10 |
0.2518 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
||||||||||||
lgb |
pyrt |
50000 |
breast_cancer |
-1 |
10 |
1 |
0.2888 |
0.2892 |
0.2889 |
0.289 |
0.289 |
0.2888 |
0.2892 |
10 |
0.289 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
lgb |
ort |
50000 |
breast_cancer |
-1 |
10 |
1 |
0.4594 |
0.4611 |
0.4602 |
0.4603 |
0.4602 |
0.4594 |
0.461 |
10 |
0.4602 |
30 |
100 |
regression |
2336 |
100 |
1218 |
2 |
1 |
|||||||||||
sklh |
1 |
digits |
-1 |
10 |
1 |
0.001929 |
0.002141 |
0.001943 |
0.001965 |
0.001971 |
0.001929 |
0.002086 |
10 |
0.001957 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
1 |
digits |
-1 |
10 |
1 |
2.169e-05 |
3.895e-05 |
2.199e-05 |
2.418e-05 |
2.444e-05 |
2.169e-05 |
3.423e-05 |
10 |
2.255e-05 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
1 |
digits |
-1 |
10 |
1 |
3.851e-05 |
9.811e-05 |
4.109e-05 |
4.811e-05 |
4.842e-05 |
3.851e-05 |
8.202e-05 |
10 |
4.144e-05 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
2 |
digits |
-1 |
10 |
1 |
0.001786 |
0.001871 |
0.001796 |
0.001817 |
0.001809 |
0.001786 |
0.001854 |
10 |
0.001801 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
2 |
digits |
-1 |
10 |
1 |
3.162e-05 |
4.258e-05 |
3.209e-05 |
3.365e-05 |
3.387e-05 |
3.162e-05 |
3.99e-05 |
10 |
3.334e-05 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
2 |
digits |
-1 |
10 |
1 |
5.185e-05 |
7.338e-05 |
5.361e-05 |
5.642e-05 |
5.625e-05 |
5.185e-05 |
6.813e-05 |
10 |
5.42e-05 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
5 |
digits |
-1 |
10 |
1 |
0.001779 |
0.001836 |
0.001794 |
0.00182 |
0.001806 |
0.001779 |
0.001836 |
10 |
0.001803 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
5 |
digits |
-1 |
10 |
1 |
5.833e-05 |
6.957e-05 |
5.976e-05 |
6.102e-05 |
6.113e-05 |
5.833e-05 |
6.729e-05 |
10 |
6.02e-05 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
5 |
digits |
-1 |
10 |
1 |
9.269e-05 |
0.000111 |
9.381e-05 |
9.675e-05 |
9.686e-05 |
9.269e-05 |
0.0001074 |
10 |
9.507e-05 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
10 |
digits |
-1 |
10 |
1 |
0.001795 |
0.001843 |
0.001805 |
0.001817 |
0.001814 |
0.001795 |
0.001843 |
10 |
0.001813 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
10 |
digits |
-1 |
10 |
1 |
0.0001011 |
0.0001126 |
0.0001034 |
0.0001058 |
0.0001048 |
0.0001011 |
0.0001109 |
10 |
0.0001036 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
10 |
digits |
-1 |
10 |
1 |
0.0001584 |
0.000178 |
0.0001609 |
0.000164 |
0.0001637 |
0.0001584 |
0.0001746 |
10 |
0.0001618 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
20 |
digits |
-1 |
10 |
1 |
0.001821 |
0.001871 |
0.001829 |
0.00184 |
0.001837 |
0.001821 |
0.001864 |
10 |
0.001837 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
20 |
digits |
-1 |
10 |
1 |
0.0001863 |
0.0002059 |
0.0001903 |
0.0001957 |
0.0001928 |
0.0001863 |
0.0002034 |
10 |
0.000191 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
20 |
digits |
-1 |
10 |
1 |
0.0002914 |
0.0003174 |
0.0002963 |
0.0003004 |
0.0002987 |
0.0002914 |
0.0003127 |
10 |
0.0002964 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
50 |
digits |
-1 |
10 |
1 |
0.00188 |
0.001921 |
0.001884 |
0.001892 |
0.00189 |
0.00188 |
0.001912 |
10 |
0.001888 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
50 |
digits |
-1 |
10 |
1 |
0.0004459 |
0.0004817 |
0.0004471 |
0.0004551 |
0.0004537 |
0.0004459 |
0.0004737 |
10 |
0.0004507 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
50 |
digits |
-1 |
10 |
1 |
0.0006944 |
0.0007288 |
0.0006983 |
0.0007025 |
0.0007025 |
0.0006944 |
0.0007207 |
10 |
0.0006999 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
100 |
digits |
-1 |
10 |
1 |
0.00198 |
0.002037 |
0.001989 |
0.002009 |
0.002001 |
0.00198 |
0.002036 |
10 |
0.001997 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
100 |
digits |
-1 |
10 |
1 |
0.0008742 |
0.0009247 |
0.0008745 |
0.0008862 |
0.0008829 |
0.0008742 |
0.0009123 |
10 |
0.0008748 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
100 |
digits |
-1 |
10 |
1 |
0.001365 |
0.0014 |
0.001371 |
0.001379 |
0.001377 |
0.001365 |
0.001397 |
10 |
0.001374 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
200 |
digits |
-1 |
10 |
1 |
0.002195 |
0.002227 |
0.002198 |
0.002212 |
0.002207 |
0.002195 |
0.002227 |
10 |
0.002207 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
200 |
digits |
-1 |
10 |
1 |
0.001725 |
0.00181 |
0.001742 |
0.001751 |
0.00175 |
0.001725 |
0.001792 |
10 |
0.001744 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
200 |
digits |
-1 |
10 |
1 |
0.002711 |
0.002774 |
0.002718 |
0.002731 |
0.002728 |
0.002711 |
0.002764 |
10 |
0.002725 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
500 |
digits |
-1 |
10 |
1 |
0.002701 |
0.002766 |
0.002723 |
0.002751 |
0.002735 |
0.002701 |
0.002766 |
10 |
0.002736 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
500 |
digits |
-1 |
10 |
1 |
0.004315 |
0.004422 |
0.004324 |
0.004337 |
0.004339 |
0.004315 |
0.004399 |
10 |
0.004329 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
500 |
digits |
-1 |
10 |
1 |
0.006551 |
0.006838 |
0.006746 |
0.006778 |
0.006728 |
0.006553 |
0.006838 |
10 |
0.006755 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
1000 |
digits |
-1 |
10 |
1 |
0.003553 |
0.003735 |
0.003562 |
0.003583 |
0.003589 |
0.003553 |
0.00369 |
10 |
0.003573 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
1000 |
digits |
-1 |
10 |
1 |
0.008602 |
0.008802 |
0.008611 |
0.008632 |
0.008636 |
0.008602 |
0.008746 |
10 |
0.008615 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
1000 |
digits |
-1 |
10 |
1 |
0.01268 |
0.01359 |
0.01272 |
0.01274 |
0.01282 |
0.01268 |
0.01334 |
10 |
0.01272 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
2000 |
digits |
-1 |
10 |
1 |
0.005061 |
0.005276 |
0.005084 |
0.005131 |
0.00512 |
0.005061 |
0.005247 |
10 |
0.005092 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
2000 |
digits |
-1 |
10 |
1 |
0.01629 |
0.01766 |
0.01665 |
0.01731 |
0.01695 |
0.01629 |
0.01766 |
10 |
0.01718 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
2000 |
digits |
-1 |
10 |
1 |
0.02545 |
0.02582 |
0.02549 |
0.02554 |
0.02553 |
0.02545 |
0.02573 |
10 |
0.02549 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
5000 |
digits |
-1 |
10 |
1 |
0.009126 |
0.009524 |
0.009153 |
0.009196 |
0.009206 |
0.009126 |
0.009438 |
10 |
0.009155 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
5000 |
digits |
-1 |
10 |
1 |
0.04079 |
0.04368 |
0.0408 |
0.04176 |
0.04143 |
0.04079 |
0.04351 |
10 |
0.0408 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
5000 |
digits |
-1 |
10 |
1 |
0.06372 |
0.06425 |
0.0638 |
0.06401 |
0.0639 |
0.06372 |
0.06422 |
10 |
0.06381 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
10000 |
digits |
-1 |
10 |
1 |
0.01587 |
0.01606 |
0.01591 |
0.01599 |
0.01594 |
0.01587 |
0.01606 |
10 |
0.01594 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
10000 |
digits |
-1 |
10 |
1 |
0.08149 |
0.08693 |
0.08159 |
0.08197 |
0.08232 |
0.08149 |
0.08539 |
10 |
0.08173 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
10000 |
digits |
-1 |
10 |
1 |
0.1273 |
0.1289 |
0.1274 |
0.1279 |
0.1278 |
0.1273 |
0.1288 |
10 |
0.1275 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
20000 |
digits |
-1 |
10 |
1 |
0.03719 |
0.03829 |
0.03734 |
0.03769 |
0.03755 |
0.03719 |
0.03817 |
10 |
0.0375 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
20000 |
digits |
-1 |
10 |
1 |
0.1631 |
0.1697 |
0.1632 |
0.1635 |
0.1639 |
0.1631 |
0.1677 |
10 |
0.1633 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
20000 |
digits |
-1 |
10 |
1 |
0.2549 |
0.2562 |
0.2552 |
0.2557 |
0.2554 |
0.2549 |
0.2561 |
10 |
0.2553 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
50000 |
digits |
-1 |
10 |
1 |
0.1179 |
0.1187 |
0.118 |
0.1183 |
0.1182 |
0.1179 |
0.1187 |
10 |
0.1182 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
|||||||||||||
sklh |
pyrt |
50000 |
digits |
-1 |
10 |
1 |
0.4076 |
0.4148 |
0.4081 |
0.4085 |
0.4088 |
0.4076 |
0.4127 |
10 |
0.4081 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
sklh |
ort |
50000 |
digits |
-1 |
10 |
1 |
0.6368 |
0.6388 |
0.6375 |
0.6382 |
0.6378 |
0.6368 |
0.6388 |
10 |
0.6378 |
0 |
0 |
64 |
100 |
2567 |
5034 |
6 |
||||||||||||
skl |
1 |
digits |
-1 |
10 |
1 |
0.00747 |
0.007993 |
0.0075 |
0.007591 |
0.00758 |
0.00747 |
0.007875 |
10 |
0.007525 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
1 |
digits |
-1 |
10 |
1 |
2.528e-05 |
6.11e-05 |
3.06e-05 |
3.794e-05 |
3.497e-05 |
2.528e-05 |
5.383e-05 |
10 |
3.28e-05 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
1 |
digits |
-1 |
10 |
1 |
4.515e-05 |
0.0001362 |
5.648e-05 |
7.555e-05 |
6.849e-05 |
4.515e-05 |
0.0001166 |
10 |
6.013e-05 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
2 |
digits |
-1 |
10 |
1 |
0.00735 |
0.007476 |
0.007378 |
0.007397 |
0.007393 |
0.00735 |
0.007473 |
10 |
0.00738 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
2 |
digits |
-1 |
10 |
1 |
3.367e-05 |
6.163e-05 |
3.686e-05 |
5.24e-05 |
4.467e-05 |
3.367e-05 |
6.163e-05 |
10 |
4.246e-05 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
2 |
digits |
-1 |
10 |
1 |
5.455e-05 |
0.0001045 |
6.029e-05 |
8.516e-05 |
7.282e-05 |
5.455e-05 |
0.0001043 |
10 |
6.748e-05 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
5 |
digits |
-1 |
10 |
1 |
0.007388 |
0.007482 |
0.007411 |
0.007439 |
0.007426 |
0.007388 |
0.007482 |
10 |
0.007423 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
5 |
digits |
-1 |
10 |
1 |
6.36e-05 |
8.644e-05 |
6.931e-05 |
8.026e-05 |
7.423e-05 |
6.36e-05 |
8.644e-05 |
10 |
7.311e-05 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
5 |
digits |
-1 |
10 |
1 |
9.56e-05 |
0.00013 |
0.0001063 |
0.0001207 |
0.0001122 |
9.56e-05 |
0.00013 |
10 |
0.0001101 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
10 |
digits |
-1 |
10 |
1 |
0.007395 |
0.007542 |
0.007415 |
0.007441 |
0.007431 |
0.007395 |
0.00751 |
10 |
0.007422 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
10 |
digits |
-1 |
10 |
1 |
0.0001091 |
0.0001356 |
0.000114 |
0.0001165 |
0.000117 |
0.0001091 |
0.0001325 |
10 |
0.0001151 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
10 |
digits |
-1 |
10 |
1 |
0.0001635 |
0.0001992 |
0.0001674 |
0.0001753 |
0.0001746 |
0.0001635 |
0.0001968 |
10 |
0.0001706 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
20 |
digits |
-1 |
10 |
1 |
0.007464 |
0.007595 |
0.007488 |
0.007521 |
0.007509 |
0.007464 |
0.007592 |
10 |
0.007494 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
20 |
digits |
-1 |
10 |
1 |
0.0002007 |
0.0002386 |
0.0002039 |
0.0002119 |
0.0002104 |
0.0002007 |
0.0002308 |
10 |
0.0002075 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
20 |
digits |
-1 |
10 |
1 |
0.0003068 |
0.0003515 |
0.0003101 |
0.0003157 |
0.0003162 |
0.0003068 |
0.000341 |
10 |
0.0003107 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
50 |
digits |
-1 |
10 |
1 |
0.007638 |
0.007771 |
0.007649 |
0.00767 |
0.007666 |
0.007638 |
0.007738 |
10 |
0.007655 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
50 |
digits |
-1 |
10 |
1 |
0.0004846 |
0.0005468 |
0.0004888 |
0.0004921 |
0.000495 |
0.0004846 |
0.0005294 |
10 |
0.0004902 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
50 |
digits |
-1 |
10 |
1 |
0.0007336 |
0.0008013 |
0.0007387 |
0.0007424 |
0.0007459 |
0.0007336 |
0.0007826 |
10 |
0.0007402 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
100 |
digits |
-1 |
10 |
1 |
0.007888 |
0.008041 |
0.007897 |
0.007919 |
0.007924 |
0.007888 |
0.008019 |
10 |
0.007899 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
100 |
digits |
-1 |
10 |
1 |
0.0009511 |
0.001038 |
0.0009574 |
0.0009672 |
0.0009682 |
0.0009511 |
0.001015 |
10 |
0.0009599 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
100 |
digits |
-1 |
10 |
1 |
0.001438 |
0.001525 |
0.001454 |
0.001461 |
0.00146 |
0.001438 |
0.001505 |
10 |
0.001456 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
200 |
digits |
-1 |
10 |
1 |
0.008507 |
0.008674 |
0.008522 |
0.008534 |
0.008541 |
0.008507 |
0.00863 |
10 |
0.008528 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
200 |
digits |
-1 |
10 |
1 |
0.00189 |
0.00201 |
0.001902 |
0.001904 |
0.001912 |
0.00189 |
0.001977 |
10 |
0.001902 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
200 |
digits |
-1 |
10 |
1 |
0.00286 |
0.002972 |
0.00287 |
0.0029 |
0.002889 |
0.00286 |
0.002952 |
10 |
0.002879 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
500 |
digits |
-1 |
10 |
1 |
0.009839 |
0.0101 |
0.009851 |
0.009889 |
0.009891 |
0.009839 |
0.01004 |
10 |
0.009868 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
500 |
digits |
-1 |
10 |
1 |
0.004723 |
0.004969 |
0.004735 |
0.004741 |
0.00476 |
0.004723 |
0.004898 |
10 |
0.004736 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
500 |
digits |
-1 |
10 |
1 |
0.007125 |
0.007511 |
0.007141 |
0.007183 |
0.007197 |
0.007125 |
0.007423 |
10 |
0.007146 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
1000 |
digits |
-1 |
10 |
1 |
0.012 |
0.01246 |
0.01204 |
0.01218 |
0.01213 |
0.012 |
0.01238 |
10 |
0.01211 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
1000 |
digits |
-1 |
10 |
1 |
0.009427 |
0.01 |
0.009459 |
0.009531 |
0.009529 |
0.009427 |
0.009846 |
10 |
0.009471 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
1000 |
digits |
-1 |
10 |
1 |
0.0142 |
0.01493 |
0.01426 |
0.0143 |
0.01434 |
0.0142 |
0.01473 |
10 |
0.01427 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
2000 |
digits |
-1 |
10 |
1 |
0.01587 |
0.0163 |
0.01598 |
0.01603 |
0.01601 |
0.01587 |
0.01622 |
10 |
0.016 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
2000 |
digits |
-1 |
10 |
1 |
0.01891 |
0.01946 |
0.01895 |
0.01897 |
0.019 |
0.01891 |
0.0193 |
10 |
0.01895 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
2000 |
digits |
-1 |
10 |
1 |
0.02854 |
0.02916 |
0.02857 |
0.02868 |
0.02866 |
0.02854 |
0.02901 |
10 |
0.0286 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
5000 |
digits |
-1 |
10 |
1 |
0.02649 |
0.0273 |
0.02668 |
0.02674 |
0.02674 |
0.02649 |
0.02714 |
10 |
0.0267 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
5000 |
digits |
-1 |
10 |
1 |
0.04738 |
0.04805 |
0.04748 |
0.04756 |
0.04755 |
0.04738 |
0.0479 |
10 |
0.04753 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
5000 |
digits |
-1 |
10 |
1 |
0.07135 |
0.0723 |
0.07142 |
0.0716 |
0.07157 |
0.07135 |
0.07209 |
10 |
0.07149 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
10000 |
digits |
-1 |
10 |
1 |
0.04415 |
0.04518 |
0.04427 |
0.04437 |
0.04439 |
0.04415 |
0.04493 |
10 |
0.04433 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
10000 |
digits |
-1 |
10 |
1 |
0.09479 |
0.09581 |
0.09489 |
0.09526 |
0.09511 |
0.09479 |
0.09577 |
10 |
0.09496 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
10000 |
digits |
-1 |
10 |
1 |
0.1427 |
0.144 |
0.143 |
0.1431 |
0.1432 |
0.1427 |
0.144 |
10 |
0.143 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
20000 |
digits |
-1 |
10 |
1 |
0.07962 |
0.08279 |
0.07972 |
0.08037 |
0.08026 |
0.07962 |
0.08201 |
10 |
0.07997 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
20000 |
digits |
-1 |
10 |
1 |
0.1895 |
0.1907 |
0.1899 |
0.1905 |
0.1901 |
0.1895 |
0.1907 |
10 |
0.1901 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
20000 |
digits |
-1 |
10 |
1 |
0.2855 |
0.2864 |
0.2859 |
0.2862 |
0.286 |
0.2855 |
0.2864 |
10 |
0.2859 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
50000 |
digits |
-1 |
10 |
1 |
0.2026 |
0.2084 |
0.2031 |
0.2041 |
0.2041 |
0.2026 |
0.2075 |
10 |
0.2035 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
|||||||||||||
skl |
pyrt |
50000 |
digits |
-1 |
10 |
1 |
0.4584 |
0.4758 |
0.4651 |
0.4752 |
0.4692 |
0.4584 |
0.4758 |
10 |
0.4747 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
skl |
ort |
50000 |
digits |
-1 |
10 |
1 |
0.7134 |
0.7149 |
0.7136 |
0.7146 |
0.7141 |
0.7134 |
0.7149 |
10 |
0.7144 |
64 |
100 |
9006 |
6 |
4553 |
64 |
1 |
||||||||||||
xgb |
1 |
digits |
-1 |
10 |
1 |
8.432e-05 |
0.0002773 |
8.535e-05 |
8.955e-05 |
0.0001065 |
8.432e-05 |
0.0002184 |
10 |
8.593e-05 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
1 |
digits |
-1 |
10 |
1 |
2.554e-05 |
5.566e-05 |
2.929e-05 |
3.24e-05 |
3.325e-05 |
2.554e-05 |
4.975e-05 |
10 |
3.123e-05 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
1 |
digits |
-1 |
10 |
1 |
4.398e-05 |
0.0001413 |
5.128e-05 |
6.055e-05 |
6.349e-05 |
4.398e-05 |
0.0001185 |
10 |
5.238e-05 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
2 |
digits |
-1 |
10 |
1 |
8.862e-05 |
0.0001322 |
8.922e-05 |
9.259e-05 |
9.456e-05 |
8.862e-05 |
0.0001195 |
10 |
8.99e-05 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
2 |
digits |
-1 |
10 |
1 |
3.299e-05 |
4.974e-05 |
3.626e-05 |
4.407e-05 |
4.018e-05 |
3.299e-05 |
4.974e-05 |
10 |
4.09e-05 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
2 |
digits |
-1 |
10 |
1 |
5.441e-05 |
7.872e-05 |
5.907e-05 |
6.858e-05 |
6.478e-05 |
5.441e-05 |
7.872e-05 |
10 |
6.65e-05 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
5 |
digits |
-1 |
10 |
1 |
0.0001066 |
0.0001275 |
0.0001069 |
0.0001112 |
0.0001106 |
0.0001066 |
0.0001227 |
10 |
0.0001082 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
5 |
digits |
-1 |
10 |
1 |
5.986e-05 |
7.349e-05 |
6.264e-05 |
6.931e-05 |
6.611e-05 |
5.986e-05 |
7.349e-05 |
10 |
6.729e-05 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
5 |
digits |
-1 |
10 |
1 |
9.602e-05 |
0.0001172 |
9.84e-05 |
0.0001086 |
0.0001036 |
9.602e-05 |
0.0001163 |
10 |
0.0001034 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
10 |
digits |
-1 |
10 |
1 |
0.0001346 |
0.0001554 |
0.0001359 |
0.0001424 |
0.0001396 |
0.0001346 |
0.0001513 |
10 |
0.0001374 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
10 |
digits |
-1 |
10 |
1 |
0.0001082 |
0.000123 |
0.0001096 |
0.0001123 |
0.0001119 |
0.0001082 |
0.0001201 |
10 |
0.0001113 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
10 |
digits |
-1 |
10 |
1 |
0.0001677 |
0.0001855 |
0.0001697 |
0.000174 |
0.0001729 |
0.0001677 |
0.0001825 |
10 |
0.0001722 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
20 |
digits |
-1 |
10 |
1 |
0.0001906 |
0.0002224 |
0.0001919 |
0.0001961 |
0.0001981 |
0.0001906 |
0.0002195 |
10 |
0.0001926 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
20 |
digits |
-1 |
10 |
1 |
0.0001942 |
0.0002191 |
0.0001999 |
0.0002029 |
0.0002022 |
0.0001942 |
0.0002148 |
10 |
0.0002006 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
20 |
digits |
-1 |
10 |
1 |
0.0003035 |
0.0003318 |
0.0003084 |
0.0003154 |
0.0003119 |
0.0003035 |
0.0003268 |
10 |
0.0003093 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
50 |
digits |
-1 |
10 |
1 |
0.0003575 |
0.0003871 |
0.0003607 |
0.0003642 |
0.0003643 |
0.0003575 |
0.0003801 |
10 |
0.0003615 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
50 |
digits |
-1 |
10 |
1 |
0.0004728 |
0.0005007 |
0.0004772 |
0.0004824 |
0.0004809 |
0.0004728 |
0.0004952 |
10 |
0.0004793 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
50 |
digits |
-1 |
10 |
1 |
0.0007322 |
0.0007584 |
0.0007386 |
0.0007454 |
0.0007422 |
0.0007322 |
0.0007562 |
10 |
0.0007418 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
100 |
digits |
-1 |
10 |
1 |
0.0006377 |
0.0006651 |
0.0006415 |
0.0006467 |
0.0006452 |
0.0006377 |
0.0006595 |
10 |
0.0006431 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
100 |
digits |
-1 |
10 |
1 |
0.0009333 |
0.0009811 |
0.0009385 |
0.0009438 |
0.0009443 |
0.0009333 |
0.0009693 |
10 |
0.0009403 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
100 |
digits |
-1 |
10 |
1 |
0.001442 |
0.001483 |
0.001446 |
0.001456 |
0.001453 |
0.001442 |
0.001475 |
10 |
0.00145 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
200 |
digits |
-1 |
10 |
1 |
0.001197 |
0.001224 |
0.0012 |
0.001207 |
0.001205 |
0.001197 |
0.00122 |
10 |
0.001204 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
200 |
digits |
-1 |
10 |
1 |
0.001852 |
0.00189 |
0.001862 |
0.001865 |
0.001865 |
0.001852 |
0.001885 |
10 |
0.001865 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
200 |
digits |
-1 |
10 |
1 |
0.002865 |
0.002958 |
0.002876 |
0.002907 |
0.002893 |
0.002865 |
0.002946 |
10 |
0.002885 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
500 |
digits |
-1 |
10 |
1 |
0.002879 |
0.002948 |
0.002888 |
0.0029 |
0.002897 |
0.002879 |
0.002935 |
10 |
0.002891 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
500 |
digits |
-1 |
10 |
1 |
0.004624 |
0.004731 |
0.004631 |
0.004637 |
0.004643 |
0.004624 |
0.004703 |
10 |
0.004632 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
500 |
digits |
-1 |
10 |
1 |
0.007126 |
0.007256 |
0.007156 |
0.007171 |
0.007167 |
0.007126 |
0.007236 |
10 |
0.007161 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
1000 |
digits |
-1 |
10 |
1 |
0.00567 |
0.005766 |
0.005689 |
0.005703 |
0.005699 |
0.00567 |
0.005747 |
10 |
0.005691 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
1000 |
digits |
-1 |
10 |
1 |
0.009231 |
0.009565 |
0.00924 |
0.00926 |
0.009279 |
0.009231 |
0.009468 |
10 |
0.009249 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
1000 |
digits |
-1 |
10 |
1 |
0.01426 |
0.01467 |
0.01429 |
0.01435 |
0.01436 |
0.01426 |
0.0146 |
10 |
0.01431 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
2000 |
digits |
-1 |
10 |
1 |
0.01125 |
0.0114 |
0.01128 |
0.01131 |
0.0113 |
0.01125 |
0.01138 |
10 |
0.01129 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
2000 |
digits |
-1 |
10 |
1 |
0.01854 |
0.01951 |
0.01858 |
0.01859 |
0.01867 |
0.01854 |
0.01922 |
10 |
0.01858 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
2000 |
digits |
-1 |
10 |
1 |
0.02852 |
0.02916 |
0.02862 |
0.02868 |
0.02868 |
0.02852 |
0.02902 |
10 |
0.02864 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
5000 |
digits |
-1 |
10 |
1 |
0.02805 |
0.02833 |
0.02808 |
0.02823 |
0.02814 |
0.02805 |
0.02833 |
10 |
0.02809 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
5000 |
digits |
-1 |
10 |
1 |
0.04644 |
0.04708 |
0.04647 |
0.0465 |
0.04655 |
0.04644 |
0.04691 |
10 |
0.04648 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
5000 |
digits |
-1 |
10 |
1 |
0.07152 |
0.07239 |
0.07164 |
0.07183 |
0.07176 |
0.07152 |
0.07223 |
10 |
0.0717 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
10000 |
digits |
-1 |
10 |
1 |
0.0559 |
0.05611 |
0.05592 |
0.05597 |
0.05596 |
0.0559 |
0.0561 |
10 |
0.05594 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
10000 |
digits |
-1 |
10 |
1 |
0.09286 |
0.09425 |
0.09295 |
0.09352 |
0.09326 |
0.09286 |
0.09407 |
10 |
0.09311 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
10000 |
digits |
-1 |
10 |
1 |
0.143 |
0.1439 |
0.1432 |
0.1434 |
0.1434 |
0.143 |
0.1438 |
10 |
0.1433 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
20000 |
digits |
-1 |
10 |
1 |
0.112 |
0.1132 |
0.112 |
0.1128 |
0.1123 |
0.112 |
0.1132 |
10 |
0.1121 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
20000 |
digits |
-1 |
10 |
1 |
0.1856 |
0.1865 |
0.1858 |
0.186 |
0.1859 |
0.1856 |
0.1864 |
10 |
0.1859 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
20000 |
digits |
-1 |
10 |
1 |
0.286 |
0.2877 |
0.2865 |
0.2873 |
0.2869 |
0.286 |
0.2877 |
10 |
0.287 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
50000 |
digits |
-1 |
10 |
1 |
0.2797 |
0.2805 |
0.28 |
0.2801 |
0.28 |
0.2797 |
0.2804 |
10 |
0.28 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
||||||||||||||
xgb |
pyrt |
50000 |
digits |
-1 |
10 |
1 |
0.4637 |
0.4656 |
0.4645 |
0.4646 |
0.4646 |
0.4637 |
0.4656 |
10 |
0.4645 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
xgb |
ort |
50000 |
digits |
-1 |
10 |
1 |
0.7156 |
0.7181 |
0.7166 |
0.7173 |
0.7168 |
0.7156 |
0.7181 |
10 |
0.7168 |
100 |
reg:linear |
9020 |
100 |
4560 |
2 |
|||||||||||||
lgb |
1 |
digits |
-1 |
10 |
1 |
0.0001056 |
0.0003095 |
0.0001094 |
0.0001217 |
0.0001331 |
0.0001056 |
0.0002494 |
10 |
0.0001111 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
1 |
digits |
-1 |
10 |
1 |
2.386e-05 |
4.973e-05 |
2.603e-05 |
3.031e-05 |
2.942e-05 |
2.386e-05 |
4.375e-05 |
10 |
2.651e-05 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
1 |
digits |
-1 |
10 |
1 |
4.002e-05 |
0.0001183 |
4.204e-05 |
5.377e-05 |
5.391e-05 |
4.002e-05 |
9.816e-05 |
10 |
4.564e-05 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
2 |
digits |
-1 |
10 |
1 |
0.0001132 |
0.000135 |
0.0001147 |
0.0001215 |
0.0001193 |
0.0001132 |
0.0001319 |
10 |
0.0001176 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
2 |
digits |
-1 |
10 |
1 |
3.278e-05 |
4.057e-05 |
3.508e-05 |
3.725e-05 |
3.588e-05 |
3.278e-05 |
4.034e-05 |
10 |
3.565e-05 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
2 |
digits |
-1 |
10 |
1 |
4.948e-05 |
6.492e-05 |
5.306e-05 |
5.627e-05 |
5.45e-05 |
4.948e-05 |
6.27e-05 |
10 |
5.365e-05 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
5 |
digits |
-1 |
10 |
1 |
0.000136 |
0.0001541 |
0.0001386 |
0.0001436 |
0.0001412 |
0.000136 |
0.0001514 |
10 |
0.0001392 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
5 |
digits |
-1 |
10 |
1 |
5.684e-05 |
7.174e-05 |
5.775e-05 |
6.044e-05 |
6.054e-05 |
5.684e-05 |
6.944e-05 |
10 |
5.911e-05 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
5 |
digits |
-1 |
10 |
1 |
8.708e-05 |
0.0001189 |
8.844e-05 |
9.839e-05 |
9.501e-05 |
8.708e-05 |
0.0001149 |
10 |
9.093e-05 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
10 |
digits |
-1 |
10 |
1 |
0.0001684 |
0.0001887 |
0.0001707 |
0.0001757 |
0.0001736 |
0.0001684 |
0.0001851 |
10 |
0.0001714 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
10 |
digits |
-1 |
10 |
1 |
9.368e-05 |
0.0001125 |
9.616e-05 |
0.0001022 |
0.0001004 |
9.368e-05 |
0.000112 |
10 |
9.874e-05 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
10 |
digits |
-1 |
10 |
1 |
0.0001459 |
0.000169 |
0.0001486 |
0.0001519 |
0.0001519 |
0.0001459 |
0.0001641 |
10 |
0.0001503 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
20 |
digits |
-1 |
10 |
1 |
0.000234 |
0.0002574 |
0.0002363 |
0.0002447 |
0.0002416 |
0.000234 |
0.000256 |
10 |
0.0002392 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
20 |
digits |
-1 |
10 |
1 |
0.0001748 |
0.0001933 |
0.0001765 |
0.0001799 |
0.0001793 |
0.0001748 |
0.0001892 |
10 |
0.0001779 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
20 |
digits |
-1 |
10 |
1 |
0.0002718 |
0.0002992 |
0.0002755 |
0.0002805 |
0.0002801 |
0.0002718 |
0.0002981 |
10 |
0.0002763 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
50 |
digits |
-1 |
10 |
1 |
0.0004242 |
0.0004511 |
0.0004283 |
0.0004336 |
0.0004319 |
0.0004242 |
0.0004463 |
10 |
0.0004294 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
50 |
digits |
-1 |
10 |
1 |
0.0004184 |
0.0005128 |
0.0004244 |
0.0004397 |
0.0004383 |
0.0004184 |
0.00049 |
10 |
0.0004291 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
50 |
digits |
-1 |
10 |
1 |
0.0006418 |
0.0007002 |
0.0006502 |
0.0006793 |
0.0006629 |
0.0006418 |
0.0007002 |
10 |
0.000652 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
100 |
digits |
-1 |
10 |
1 |
0.0007459 |
0.0007774 |
0.0007509 |
0.0007751 |
0.0007578 |
0.0007459 |
0.0007774 |
10 |
0.0007524 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
100 |
digits |
-1 |
10 |
1 |
0.0008123 |
0.000846 |
0.0008152 |
0.0008328 |
0.0008229 |
0.0008123 |
0.0008439 |
10 |
0.0008189 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
100 |
digits |
-1 |
10 |
1 |
0.00127 |
0.001309 |
0.001271 |
0.001276 |
0.001277 |
0.00127 |
0.001298 |
10 |
0.001273 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
200 |
digits |
-1 |
10 |
1 |
0.001383 |
0.001424 |
0.001389 |
0.0014 |
0.001396 |
0.001383 |
0.001418 |
10 |
0.001391 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
200 |
digits |
-1 |
10 |
1 |
0.001607 |
0.001636 |
0.001616 |
0.001623 |
0.00162 |
0.001607 |
0.001636 |
10 |
0.001617 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
200 |
digits |
-1 |
10 |
1 |
0.002517 |
0.002572 |
0.002523 |
0.002537 |
0.002531 |
0.002517 |
0.002561 |
10 |
0.002525 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
500 |
digits |
-1 |
10 |
1 |
0.003297 |
0.003339 |
0.003304 |
0.003311 |
0.003309 |
0.003297 |
0.003331 |
10 |
0.003305 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
500 |
digits |
-1 |
10 |
1 |
0.003998 |
0.004069 |
0.004008 |
0.004022 |
0.004017 |
0.003998 |
0.004055 |
10 |
0.004011 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
500 |
digits |
-1 |
10 |
1 |
0.00625 |
0.006355 |
0.00627 |
0.006281 |
0.006278 |
0.00625 |
0.006332 |
10 |
0.006274 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
1000 |
digits |
-1 |
10 |
1 |
0.006477 |
0.00658 |
0.006494 |
0.00651 |
0.006511 |
0.006477 |
0.006564 |
10 |
0.006508 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
1000 |
digits |
-1 |
10 |
1 |
0.007993 |
0.008152 |
0.008011 |
0.008028 |
0.008027 |
0.007993 |
0.008113 |
10 |
0.008019 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
1000 |
digits |
-1 |
10 |
1 |
0.01248 |
0.01269 |
0.0125 |
0.01252 |
0.01252 |
0.01248 |
0.01263 |
10 |
0.01251 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
2000 |
digits |
-1 |
10 |
1 |
0.01287 |
0.01302 |
0.01289 |
0.01293 |
0.01291 |
0.01287 |
0.013 |
10 |
0.0129 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
2000 |
digits |
-1 |
10 |
1 |
0.01601 |
0.01632 |
0.01602 |
0.01606 |
0.01606 |
0.01601 |
0.01623 |
10 |
0.01604 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
2000 |
digits |
-1 |
10 |
1 |
0.02496 |
0.02535 |
0.02501 |
0.0251 |
0.02508 |
0.02496 |
0.02534 |
10 |
0.02503 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
5000 |
digits |
-1 |
10 |
1 |
0.03212 |
0.03245 |
0.03229 |
0.03233 |
0.03227 |
0.03212 |
0.03245 |
10 |
0.03229 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
5000 |
digits |
-1 |
10 |
1 |
0.04012 |
0.04051 |
0.04017 |
0.04025 |
0.04023 |
0.04012 |
0.04044 |
10 |
0.04019 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
5000 |
digits |
-1 |
10 |
1 |
0.06271 |
0.06321 |
0.06282 |
0.06289 |
0.06288 |
0.06271 |
0.06318 |
10 |
0.06283 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
10000 |
digits |
-1 |
10 |
1 |
0.06401 |
0.06467 |
0.06414 |
0.06427 |
0.06422 |
0.06401 |
0.06455 |
10 |
0.06419 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
10000 |
digits |
-1 |
10 |
1 |
0.08019 |
0.08077 |
0.08031 |
0.08037 |
0.08037 |
0.08019 |
0.0807 |
10 |
0.08034 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
10000 |
digits |
-1 |
10 |
1 |
0.1252 |
0.1259 |
0.1255 |
0.1259 |
0.1256 |
0.1252 |
0.1259 |
10 |
0.1257 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
20000 |
digits |
-1 |
10 |
1 |
0.1283 |
0.1287 |
0.1284 |
0.1285 |
0.1284 |
0.1283 |
0.1287 |
10 |
0.1284 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
20000 |
digits |
-1 |
10 |
1 |
0.1605 |
0.161 |
0.1606 |
0.1609 |
0.1607 |
0.1605 |
0.161 |
10 |
0.1607 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
20000 |
digits |
-1 |
10 |
1 |
0.2505 |
0.2521 |
0.2511 |
0.2517 |
0.2513 |
0.2505 |
0.2521 |
10 |
0.2512 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
50000 |
digits |
-1 |
10 |
1 |
0.3204 |
0.3235 |
0.321 |
0.3227 |
0.3217 |
0.3204 |
0.3235 |
10 |
0.3216 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
||||||||||||
lgb |
pyrt |
50000 |
digits |
-1 |
10 |
1 |
0.4016 |
0.4028 |
0.4019 |
0.4025 |
0.4022 |
0.4016 |
0.4028 |
10 |
0.4022 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
|||||||||||
lgb |
ort |
50000 |
digits |
-1 |
10 |
1 |
0.6265 |
0.6294 |
0.6278 |
0.6289 |
0.6281 |
0.6265 |
0.6294 |
10 |
0.628 |
64 |
100 |
regression |
4974 |
100 |
2537 |
2 |
1 |
Benchmark code#
# coding: utf-8
"""
Benchmark of a couple of machine learning frameworks
for random forest.
"""
# Authors: Xavier Dupré (benchmark)
# License: MIT
import matplotlib
matplotlib.use('Agg')
import os
from logging import getLogger
from time import perf_counter as time
import numpy
import pandas
import matplotlib.pyplot as plt
import sklearn
from sklearn.experimental import enable_hist_gradient_boosting
from sklearn.ensemble import HistGradientBoostingRegressor, RandomForestRegressor
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from sklearn.utils._testing import ignore_warnings
from sklearn.datasets import load_breast_cancer, load_digits, make_classification
from xgboost import XGBRegressor
from lightgbm import LGBMRegressor
from mlprodict.onnxrt import OnnxInference
from pymlbenchmark.context import machine_information
from pymlbenchmark.benchmark import BenchPerf, BenchPerfTest
from pymlbenchmark.plotting import plot_bench_results, plot_bench_xtime
from skl2onnx import to_onnx
from onnxruntime import InferenceSession
from mlprodict.onnx_conv import register_converters
from mlprodict.tools.model_info import analyze_model
filename = os.path.splitext(os.path.split(__file__)[-1])[0]
def create_datasets():
results = {}
X, y = load_breast_cancer(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y)
results['breast_cancer'] = [X_train, X_test, y_train, y_test]
X, y = load_digits(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=41)
results['digits'] = [X_train, X_test, y_train, y_test]
X, y = make_classification(20000, 20)
X_train, X_test, y_train, y_test = train_test_split(X, y)
results['rndbin100'] = [X_train, X_test, y_train, y_test]
return results
register_converters()
common_datasets = create_datasets()
def get_model(lib):
if lib == "sklh":
return HistGradientBoostingRegressor(max_depth=6, max_iter=100)
if lib == "skl":
return RandomForestRegressor(max_depth=6, n_estimators=100)
if lib == 'xgb':
return XGBRegressor(max_depth=6, n_estimators=100)
if lib == 'lgb':
return LGBMRegressor(max_depth=6, n_estimators=100)
raise ValueError("Unknown library '{}'.".format(lib))
class LibOrtBenchPerfTest(BenchPerfTest):
def __init__(self, lib, dataset):
BenchPerfTest.__init__(self)
logger = getLogger("skl2onnx")
logger.propagate = False
logger.disabled = True
self.dataset_name = dataset
self.lib_name = lib
self.models = {}
self.datas = {}
self.onxs = {}
self.orts = {}
self.oinfcs = {}
self.model_info = {}
self.output_name = {}
self.input_name = {}
self.models[lib] = get_model(lib)
self.datas[lib] = common_datasets[dataset]
x = self.datas[lib][0]
y = self.datas[lib][2]
self.models[lib].fit(x, y)
self.model_info[lib] = analyze_model(self.models[lib])
try:
self.onxs[lib] = to_onnx(
self.models[lib],
self.datas[lib][0][:1].astype(numpy.float32),
options=None)
except RuntimeError:
pass
if lib in self.onxs:
self.orts[lib] = InferenceSession(
self.onxs[lib].SerializeToString())
self.oinfcs[lib] = OnnxInference(
self.onxs[lib], runtime='python_compiled')
self.output_name[lib] = self.oinfcs[lib].output_names[-1]
self.input_name[lib] = self.orts[lib].get_inputs()[0].name
def fcts(self, **kwargs):
def predict_ort(X, model, namei):
try:
return model.run(None, {namei: X})[1]
except Exception as e:
return None
def predict_model(X, model):
return model.predict(X)
def predict_pyrtc(X, model, namei, nameo):
return model.run({namei: X})[nameo]
res = []
for lib in [self.lib_name]:
res.append({'lib': lib, 'rt': '',
'fct': lambda X: predict_model(X, self.models[lib])})
if lib in self.oinfcs:
res.append({'lib': lib, 'rt': 'pyrt',
'fct': lambda X: predict_pyrtc(
X, self.oinfcs[lib], self.input_name[lib],
self.output_name[lib])})
if lib in self.orts:
res.append({'lib': lib, 'rt': 'ort',
'fct': lambda X: predict_ort(
X, self.orts[lib], self.input_name[lib])})
return res
def data(self, N=10, dim=-1, **kwargs): # pylint: disable=W0221
if dim != -1:
raise ValueError("dim must be -1 as it is fixed.")
lib = self.lib_name
x = self.datas[lib][1]
nbs = numpy.random.randint(0, x.shape[0] - 1, N)
res = x[nbs, :].astype(numpy.float32)
return (res, )
def validate(self, results, **kwargs):
final = {}
for k, v in self.model_info[self.lib_name].items():
final['fit_' + k] = v
return final
@ignore_warnings(category=FutureWarning)
def run_bench(repeat=10, verbose=False):
pbefore = dict(dim=[-1],
lib=['sklh', 'skl', 'xgb', 'lgb'],
dataset=["breast_cancer", "digits", "rndbin100"])
pafter = dict(N=[1, 2, 5, 10, 20, 50, 100, 200, 500, 1000,
2000, 5000, 10000, 20000, 50000])
test = lambda dim=None, **opts: LibOrtBenchPerfTest(**opts)
bp = BenchPerf(pbefore, pafter, test)
with sklearn.config_context(assume_finite=True):
start = time()
results = list(bp.enumerate_run_benchs(repeat=repeat, verbose=verbose,
stop_if_error=False))
end = time()
results_df = pandas.DataFrame(results)
print("Total time = %0.3f sec\n" % (end - start))
return results_df
if True:
#########################
# Runs the benchmark
# ++++++++++++++++++
df = run_bench(verbose=True)
df.to_csv("%s.perf.csv" % filename, index=False)
print(df.head())
#########################
# Extract information about the machine used
# ++++++++++++++++++++++++++++++++++++++++++
pkgs = ['numpy', 'pandas', 'sklearn', 'skl2onnx',
'onnxruntime', 'onnx', 'mlprodict']
dfi = pandas.DataFrame(machine_information(pkgs))
dfi.to_csv("%s.time.csv" % filename, index=False)
print(dfi)
else:
df = pandas.read_csv("%s.perf.csv" % filename)
print(df.head())
#############################
# Plot the results
# ++++++++++++++++
def label_fct(la):
la = la.replace("-lib=", "")
la = la.replace("rt=", "-")
return la
plot_bench_results(df, row_cols=('rt',), col_cols=('dataset', ), label_fct=label_fct,
x_value='N', hue_cols=('lib',), cmp_col_values='lib',
title="Numerical datasets\nBenchmark scikit-learn, xgboost, lightgbm")
plt.savefig("%s.curve.png" % filename)
import sys
if "--quiet" not in sys.argv:
plt.show()